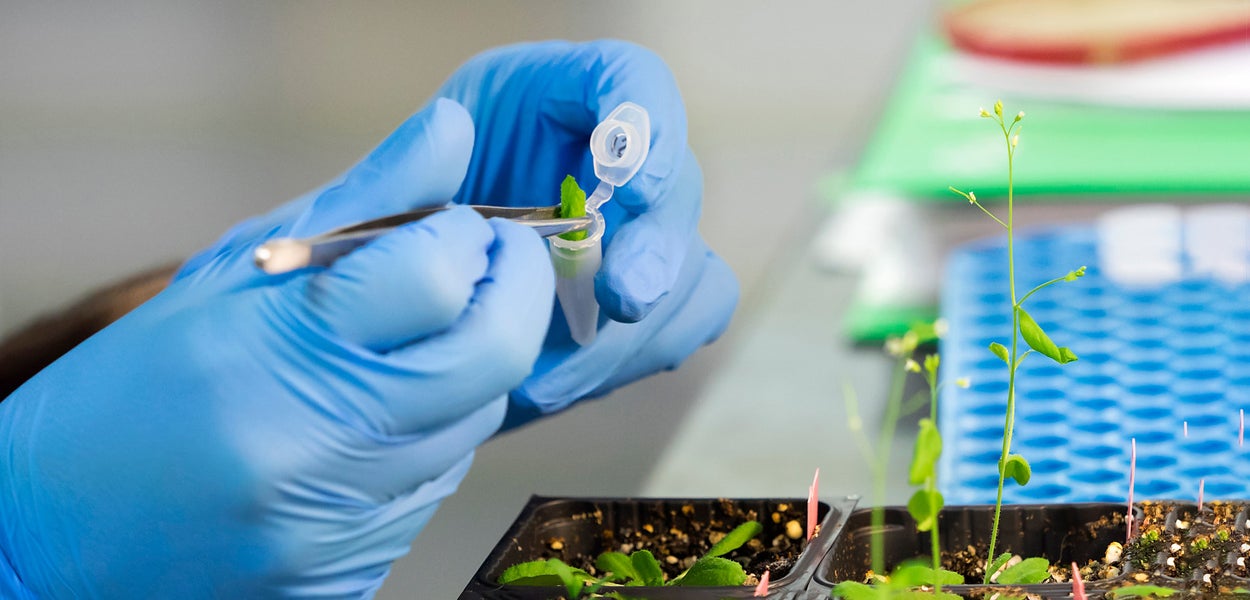
AIMMS understands molecules, makes molecules, detects molecules and studies their impact in biotechnological, biomedical and environmental contexts.
The mission of AIMMS is to achieve breakthroughs in molecular, pharmaceutical and life sciences, through fundamental understanding of molecules and their interaction with biological systems, humans and the environment. AIMMS integrates these disciplines to understand, modulate and use biological processes to improve sustainability and human health.